Individual-based modelling
The term individual-based modelling describes a group of simulation, in which the local rules of interaction between elements of the system are specified, and then an evolution of the system consisting of many such elements is investigated by computer simulation. Using this approach, a complex global behaviour of a model is obtained as a result of many simultaneous, distributed in space, local and simple interactions.
Individual-based models are natural for many physical, chemical or biological systems, whose behaviour is a result of interactions of many active elements distributed in space. They could be especially beneficial when, besides a mere numerical agreement with a physical system, the model should map its structure and topology. For such systems, individual-based modelling can be an attractive substitute for models based on differential equations apparatus. Moreover individual-based approach makes it possible to work with models of complex biological systems, which would be impossible even to formulate in traditional mathematical frameworks. Cellular automata, lattice gas automata, molecular dynamics and particle models are well known formal context for individual-based modelling.
DigiHive environment
DigiHive environment is an original, artificial, complete, low level, and closed environment consisting of space, moving objects, and rules which govern their interactions. The environment operates on two levels. On the first level, entities move and collide. Collisions result in rebounding off entities or randomly changing their internal structures. On the second level some structures of entities are capable of inducing specific changes in other entities. Types of the changes are encoded in structures of entities in simplified Prolog language. The simulation can start from any initial configuration of entities thus allowing to model various systems.
Main window
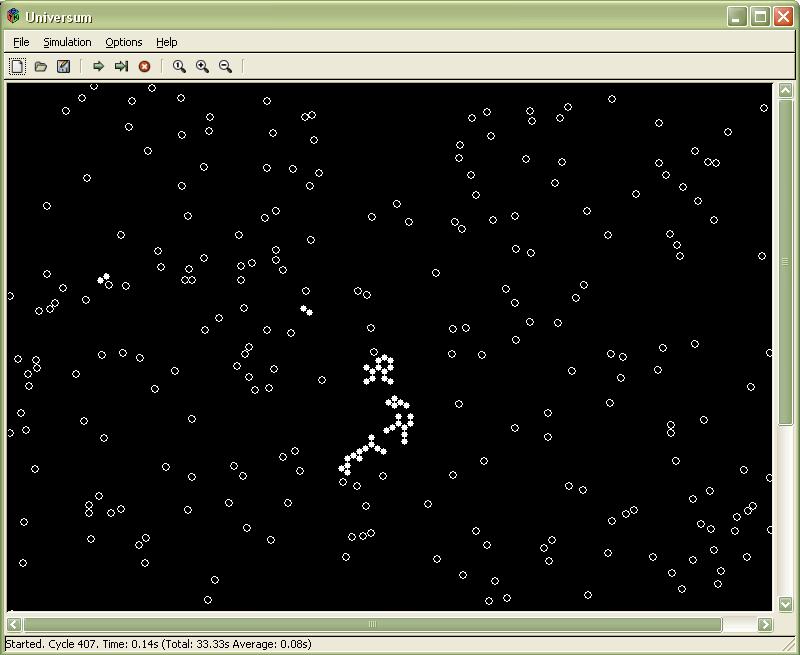
Circles represent basic constituent objects called particles. Particles move and collide according to rules of a simplified classical mechanics (including the conservation of energy and the conservation of momentum laws). Particles can bound together forming a spatial structure called complex of particles. Bound particles (part of complex) are showed as filled circles.
In the figure a sample simulation is running – state after 407 time cycles. Last step took 0.14s, the whole simulation took 33.33s, an average time step took 0.08s. Clicking on the particle opens Complex Detail window described below.
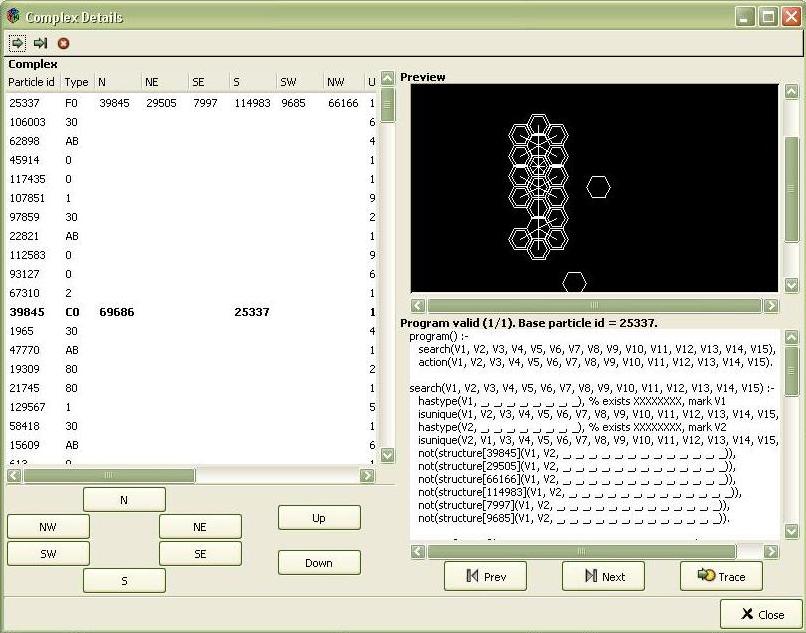
Complex details
The screenshot presents the detailed view of a particle and its neighbourhood. The window is divided into 3 section: Preview, Complex and Programs. Preview section contains a magnified view of the selected particle and its neighbourhood. The horizontal bonds are drawn using lines, particles with vertical bonds are drawn using double lines. Complex section contains a list of all the particles forming the complex to which the selected particle belongs. All physical attributes of a particle are shown. The attributes belonging to the selected particle are printed in bold. The navigation buttons in the lower part of the section change the selected particle (moves the asterisk in Preview section in the direction according to the label on the button). Programs section lists all the programs contained in the complex to which the selected particle belongs. The buttons Prev and Next navigate through the list. Trace button opens a simple debugger, described below.
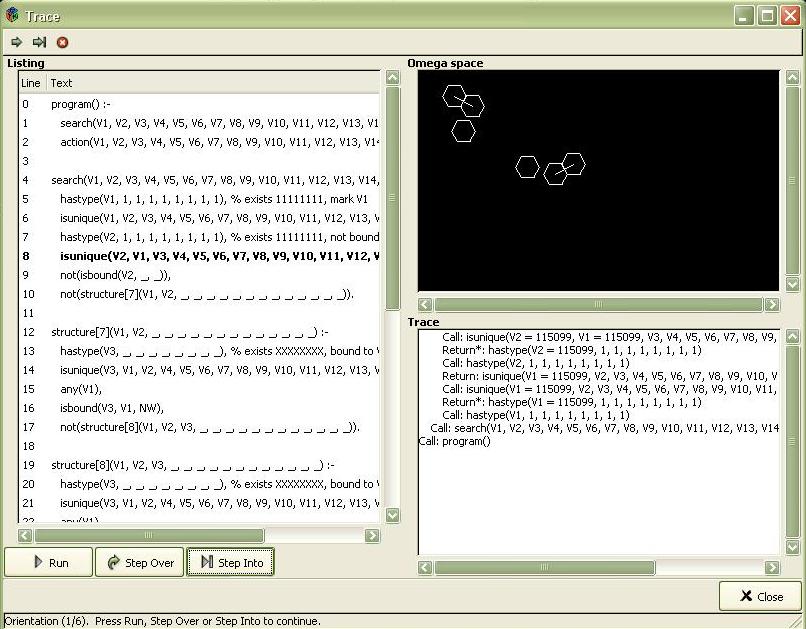
Debugger
Debugger window consists of 3 sections: Listing, Omega Space and Trace. Listing section shows a listing of the programs being debugged. The currently executed command is printed with the bold font. The lower part of the section contains the following buttons:
- Run: runs the currently displayed program,
- Step Into: runs exactly one statement of the currently displayed program. If the statement is a call to a clause, the next statement displayed is the first statement in the clause.
- Step Over: runs one command of the currently displayed program. If the statement is a call to a clause, the next statement displayed is the first statement of the current clause.